(I) Taxol treatment, which prevents microtubule depolymerization, arrests the cell at this stage.
(II) With a mitogen treatment such as an epidermal growth factor and arrested cell at this stage proceed to the next stage of the cell cycle.
(III) The cell cycle checkpoint at this stage confirms that DNA duplication is complete before the cell proceeds to the next stage.
Know your College Admission Chances Based on your Rank/Percentile, Category and Home State.
Get your JEE Main Personalised Report with Top Predicted Colleges in JoSA
This question involves identifying stages of the cell cycle (G1, S, G2, M) from graphs showing DNA content per cell and matching them with specific cellular activities or treatments.
The cell cycle consists of:
- G1 phase: Cell growth before DNA replication. DNA content is 2C (diploid).
- S phase: DNA synthesis (replication). DNA content increases from 2C to 4C.
- G2 phase: Preparation for mitosis. DNA content is 4C.
- M phase: Mitosis (cell division). DNA content decreases from 4C to 2C after cytokinesis.
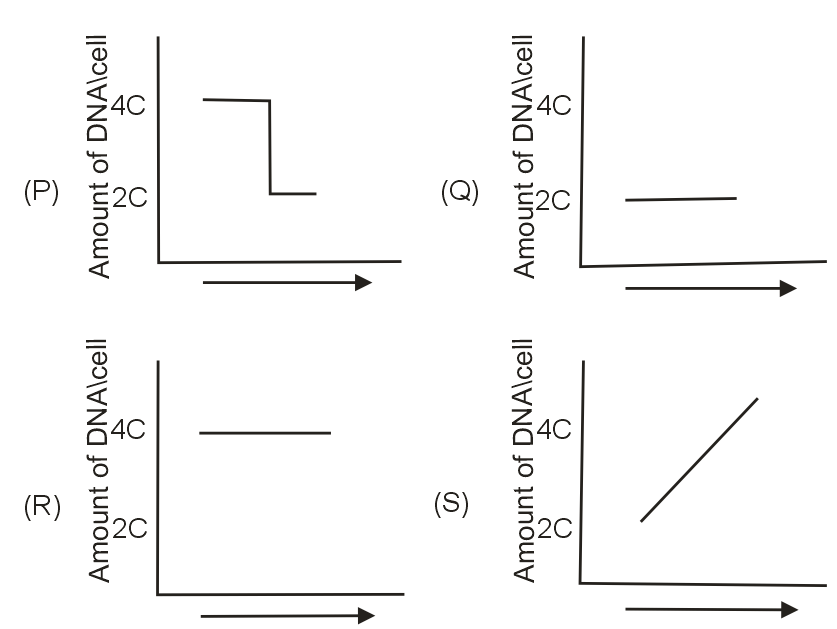
Analyzing the graphs (P, Q, R, S) based on DNA content:
- Graph with lowest DNA content (2C): G1 phase
- Graph with intermediate DNA content (between 2C-4C): S phase
- Graph with highest DNA content (4C): G2 or M phase (before division)
- Graph showing decrease from 4C to 2C: M phase (during division)
Now let's analyze the three statements:
(I) Taxol treatment: Taxol prevents microtubule depolymerization, which is essential for spindle formation during mitosis. This arrests cells in M phase.
(II) Mitogen treatment: Mitogens like epidermal growth factor stimulate cells to exit G0 (quiescence) and enter G1 phase, allowing them to proceed through the cell cycle.
(III) DNA duplication checkpoint: This occurs at the G2/M checkpoint which verifies that DNA replication is complete before entering mitosis. This corresponds to G2 phase.
Now matching the graphs to stages:
- Graph P: Shows cells with 2C DNA content → G1 phase
- Graph Q: Shows cells with DNA content between 2C-4C → S phase
- Graph R: Shows cells with 4C DNA content → G2 phase
- Graph S: Shows cells with decreasing DNA content (4C→2C) → M phase
Matching the activities:
- (I) Taxol arrests at M phase → Graph S
- (II) Mitogen treatment affects G1 phase → Graph P
- (III) DNA duplication checkpoint at G2 phase → Graph R
Therefore, the correct matching is: I - S, II - P, III - R
Looking at the options provided:
- I - P, II - S, III - Q → Incorrect
- I - R, II - Q, III - S → Incorrect
- I - P, II - Q, III - R → Incorrect
- I - Q, II - S, III - R → Incorrect
Note: None of the given options exactly match our analysis (I-S, II-P, III-R). However, based on the question and standard cell cycle knowledge, this is the correct identification.
Related Topics
Cell Cycle Checkpoints: Critical control points that ensure proper progression through the cell cycle. The main checkpoints are at G1/S, G2/M, and during mitosis.
Mitotic Spindle: Structure composed of microtubules that separates chromosomes during mitosis. Drugs like taxol stabilize microtubules, preventing proper spindle function.
Growth Factors: Signaling molecules (like epidermal growth factor) that stimulate cell division by promoting entry into G1 phase from G0.
Key Formulae
DNA content changes during cell cycle: